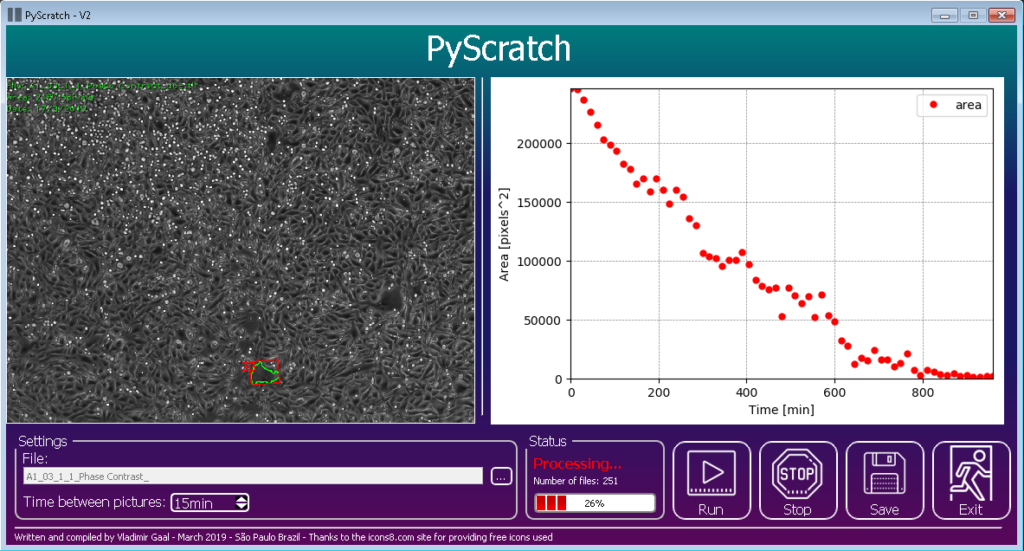
PyScratch is a new open-source software implemented in Python for migration data assays analysis, with a user-friendly interface allowing scientists with a few or none programming skills to use it.
The software was designed in a partnership between three Brazilian scientists in the NanoCell Interactions lab at the University of Campinas in 2017.
It was born from the necessity of these scientists, and today the software’s purpose is to facilitate the daily practice of researchers, exclude the manual analysis, increase reproducibility, and minimize human errors.
Image analysis has been one of the most important ways used by scientists in different methodologies to analyze results.
Not only the use of automated microscopes has increased in the past years, but also the complexity of the data acquired.
Part of a scientist’s job is to discover how to deal with a new kind of information, besides, analyze and process data.
For that to happen, scientists need good and specialized tools to correctly extract and interpret all the data.
PyScratch was first created to help Fernanda Garcia-Fossa, a biologist researcher, to analyze a huge amount of data from her migration assays.
“I performed the cancer cells’ scratch and incubated in the equipment for 48 hours, acquiring pictures every 15 minutes, and at the end of only one experiment, I had about a thousand pictures to look at and analyze!
It was impossible to do it manually”, says Garcia-Fossa. To solve that problem Garcia-Fossa went to her partner, the physics Vladimir Gaal to help, in that time Gaal was learning Python, which was a great opportunity to put that knowledge into practice.
So, both of them worked in the solution through a Python routine, which recognized the scratched areas and exported to a csv file.
“With the time, we felt the necessity to develop a user interface making even easier to use, in that way we could publish the software for any researcher that needed to use as well”, says Garcia-Fossa about the software article that you can check it out clicking here.
Garcia-Fossa also tells that they took a while to recognize and define the migration area from the images since the pictures can be very different one from another because of light, focus and contrast, and the version used today can already analyze pretty well.
Nevertheless, they are still working on the software launching new and better versions, since the published article has brought up some improvement needs because of the users’ demand.
The assay used to validate the software performance, the migration assay, or scratch assay, or wound healing, is an assay commonly used in biology because it turns possible to analyze the underlying mechanism of physiological and pathological cellular events.
Study wound healing is an important way to understand the development and tissue modeling, besides angiogenesis and tumor development.
When the assay is performed in two dimensions, it is possible to measure how fast cells react to the wound, covering the defined area.
In another words, the experiment basically is, create a gap in the monolayer of confluence cells in a plaque.
Over time, to fill the gap the cells start to migrate, and the velocity of cells migration can be measured.
Then, to measure the cell migration velocity they need to acquire images, a bunch of it, which in turn is a problematic step of the analysis because it requires manual measurement.
Fortunately, today we have at our disposal a lot of technology available to improve and upgrade our pipeline of analysis, allowing scientists to adopt better and more personalized ways to get a result.
Like Garcia-Fossa and Gaal did it.
Today is possible to find other commercial and non-commercial tools available to process the wound area.
But they are not simply as PyScratch, and require from the user some level of programming and also require full-time attention from the user, making the analysis more susceptible to human errors, plus the time the researcher takes to analyze all the pictures and data.
In the article, the authors explain how the software works. From all the pictures made in the experiment, the user gets a comma-separated values (.cvs) file, an output that stores tabular data in plain text.
The user then can process the data in their usual routine. Garcia-Fossa says that the program was essential, for her master’s thesis, “The software transforms the input data into value that make biological sense, like cell migration velocity.
I was able to better analyze the effect of my nanoparticle on prostate cancer cells and measure the cell migration velocity, and the exact time for the good closure, all thanks to PyScratch.”
If you want to try PyScratch for your research, the software is freely available and everyone from the scientific community is allowed to use it, helping Garcia-Fossa and Gaal to enhance and improve the program.
____
How is your Mind the Graph experience going so far? Help us to improve our platform for you and for a lot of other scientists posting a Mind the Graph review. Tell us about your experience, just click here.

Subscribe to our newsletter
Exclusive high quality content about effective visual
communication in science.